The crowded and confined environment inside living cells poses major challenges for high-resolution structural characterization of biomacromolecules. Site-specific protein labeling with functional probes or tags is an effective approach for characterizing protein structure, interactions, and dynamic behavior. Although substantial progress has been made in the field of protein modification, existing protein-labeling strategies still fall short of meeting the biophysical requirements for high-resolution studies of the motions of proteins and other biomacromolecules in living cells. Because of its high detection sensitivity and the absence of endogenous cellular background signals, 19F-NMR has shown great potential for protein analysis in living cells. However, current 19F-labeling methods suffer from problems such as the low efficiency of incorporating fluorinated amino acids and signal heterogeneity. Moreover, the introduction of unnatural fluorinated amino acids through genetic code expansion may lead to cytotoxicity and incompatibility with tRNA synthetases.
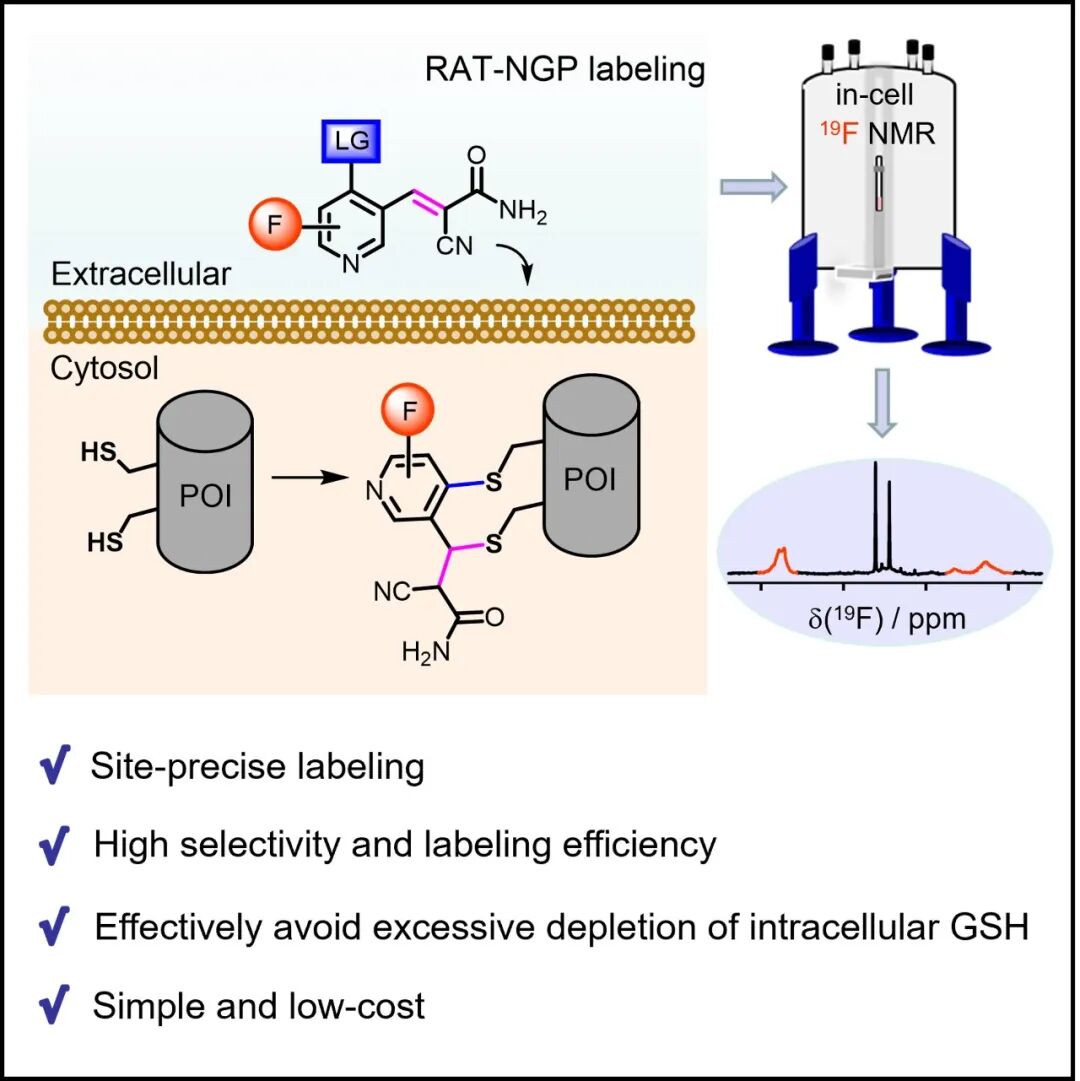
Recently, the research group of Professor Xuncheng Su at Nankai University developed a site-specific protein-labeling method in living cells based on a “reversible addition reaction-triggered neighboring-group participation” (RAT-NGP) chemical reaction. This labeling strategy is based on a reactive small-molecule tag that undergoes two sequential reactions with two engineered, solvent-accessible cysteine residues on the target protein (POI-2C). Each functional tag molecule contains two reactive groups: one is a Michael acceptor that reacts reversibly with a free thiol, and the other is a leaving group (LG) that reacts selectively with thiols. Compared with the reversible Michael acceptor, the leaving group is less reactive toward free thiols, and its reactivity can be tuned. In POI-2C, the thiol side chains of the two cysteine residues are separated by a certain spatial distance, typically about 5–7 Å. First, one free thiol is linked to the tag molecule through a reversible Michael addition. This first-step reaction brings the leaving group (LG) in the tag into closer proximity to the other thiol in the protein, thereby promoting the nucleophilic substitution of the LG by that thiol. The second step is analogous to a neighboring-group participation (NGP) reaction in organic synthesis. Within an appropriate spatial range, the two protein thiols become linked to the tag through a cascade reaction, thereby achieving site-specific labeling of the target protein. The distinctive feature of this strategy is that the reversible Michael addition triggers the NGP effect, while the NGP reaction further stabilizes the reversible addition product. At the same time, it effectively avoids interference from a single free thiol, thereby enabling site-specific labeling of the target protein. Relevant achievements were published in CCS Chemistry, 2026, DOI: 10.31635/ccschem.025.202506343.